Kinetic Accessibility of Buried DNA Sites in Nucleosomes
Wolfram Möbius, Richard A. Neher and Ulrich Gerland
With help of a light microscope one can see single chromosomes, the closest packed form of DNA. Not only during cell division, but throughout the whole cell cycle, DNA in eucaryotes is packed in one way or another. On the smallest scale, DNA is wrapped around histones and forms nucleosomes. The readout of information - between cell divisions - is controlled by so called transcription factors, which can bind to DNA without ATP consumption. How can that be if DNA is packed? On the scale of a single nucleosome, this paradoxon can be understood in the context of the so called site exposure mechanism (Polach and Widom, 1995). Thermal fluctuation partially unwrap the DNA from the histone complex making DNA accessible for transcription factor binding. For biological functioning it is not only important how DNA is made accessible, but also on which time scale this happens. Two experiments in vitro (Li et al., 2005, and Tomschik et al., 2005) have been done recently to study this question.
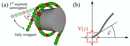
We studied the problem using a theoretical model, where the DNA was represented by beads and springs behaving like a worm-like chain. The interaction with the histone complex was mimiced by attractive potentials resulting in DNA to form a superhelix. The figure (a) illustrates the geometry and how unwrapping of the DNA looks like. Having noticed that within this model, DNA wraps and unwraps in segments of about 10 base pairs, we focused on the functional dependence of the wrapping/unwrapping rates for site exposure and re-burial on the site position. Close inspection revealed a surprising physical effect of flexibility-assisted barrier crossing, which we characterized within a toy model, the Semiflexible Brownian Rotor as sketched in figure (b).